

A high-throughput assay for profiling protein-peptide interactions
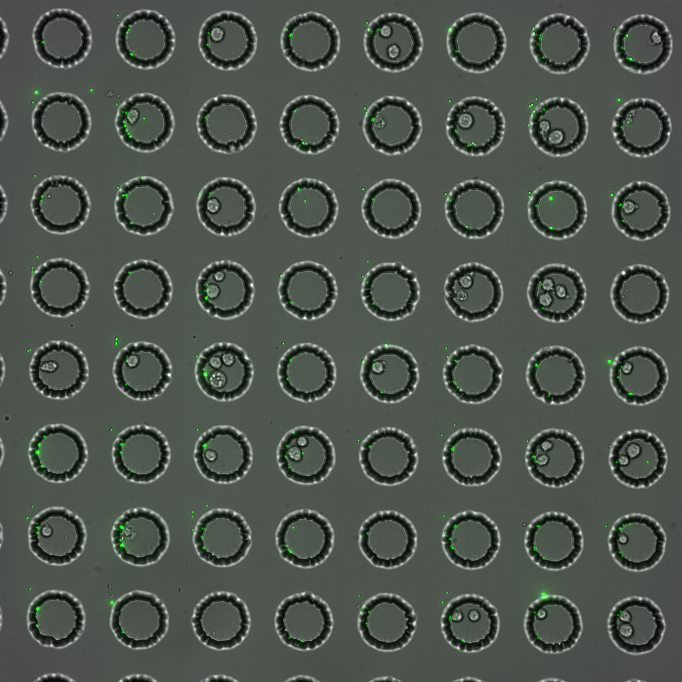
While advances in deep sequencing have dramatically improved our ability to profile how DNA and RNA binding proteins interact with their substrates, we lack similar tools for quantitative, high-throughput investigation of protein-peptide interactions. We have recently demonstrated that peptides can be synthesized directly on our encoded MRBLEs, laying the foundation for novel high-throughput on-bead assays to probe how proteins recognize their peptide substrates. In collaboration with the Cyert Lab, we are using this platform to understand how calcineurin, a human phosphatase essential for the immune response and the target of several immunosuppresents, binds its targets.
A cheap and scalable way to synthesize custom peptide libraries on encoded beads
The ability to flexible construct specific peptide libraries on spectrally encoded beads would provide a powerful proteomic tool. Using a commercial video projector ($300, Amazon.com), we are developing a new system that uses patterned light to synthesize specific peptide sequences onto beads spatially arrayed within a microfluidic device.
Using encoded beads to link genotype and phenotype for many single cells in parallel
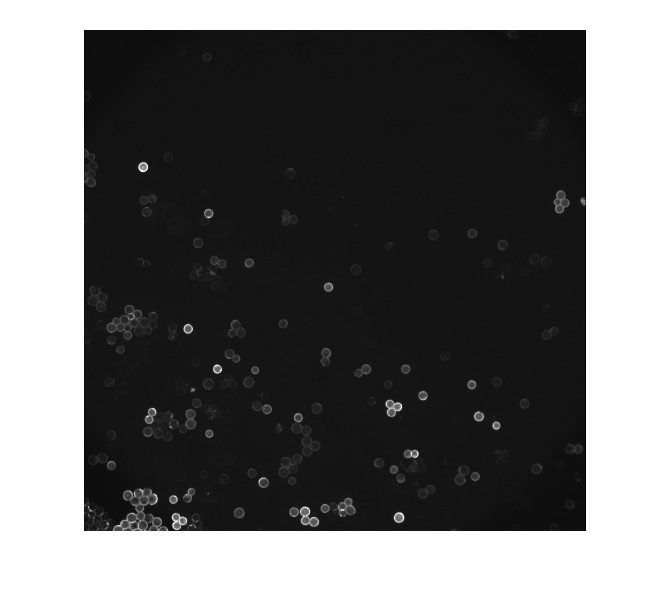
Recent developments in single-cell analysis have enabled the probing of phenotypes, transcriptomes, and genomes for individual cells, revealing previously hidden heterogeneity within cell populations of high significance for human health and disease. The next great challenge is to understand both the cause and effect of this heterogeneity by linking changes in genetic information with their functional effects. We are developing a new technologies that uses oligonucleotide arrays coupled to encoding beads to link different types of measurements for the same cell.